For decades, the field of vaccinology has been engaged in a high-stakes arms race against some of the most formidable pathogens in human history. Viruses like HIV, Ebola, and SARS-CoV-2 are masterfully adapted to evade the human immune system, often utilizing sophisticated surface proteins to gain entry into cells. While these proteins serve as the primary targets for neutralizing antibodies, studying them has long been hampered by a technical paradox: in order to analyze these proteins in the laboratory, scientists have traditionally been forced to strip them of the very environment that defines their behavior.
A breakthrough study published in Nature Communications by a multi-disciplinary team at Scripps Research, in partnership with IAVI and other global collaborators, has introduced a transformative solution. By utilizing "nanodisc" technology, the researchers have developed a platform that embeds viral proteins into lipid-based structures, effectively recreating the native viral membrane. This innovation allows scientists to observe how antibodies interact with viruses in a context that is finally, accurately, human-relevant.
The Structural Dilemma: Why Conventional Methods Fall Short
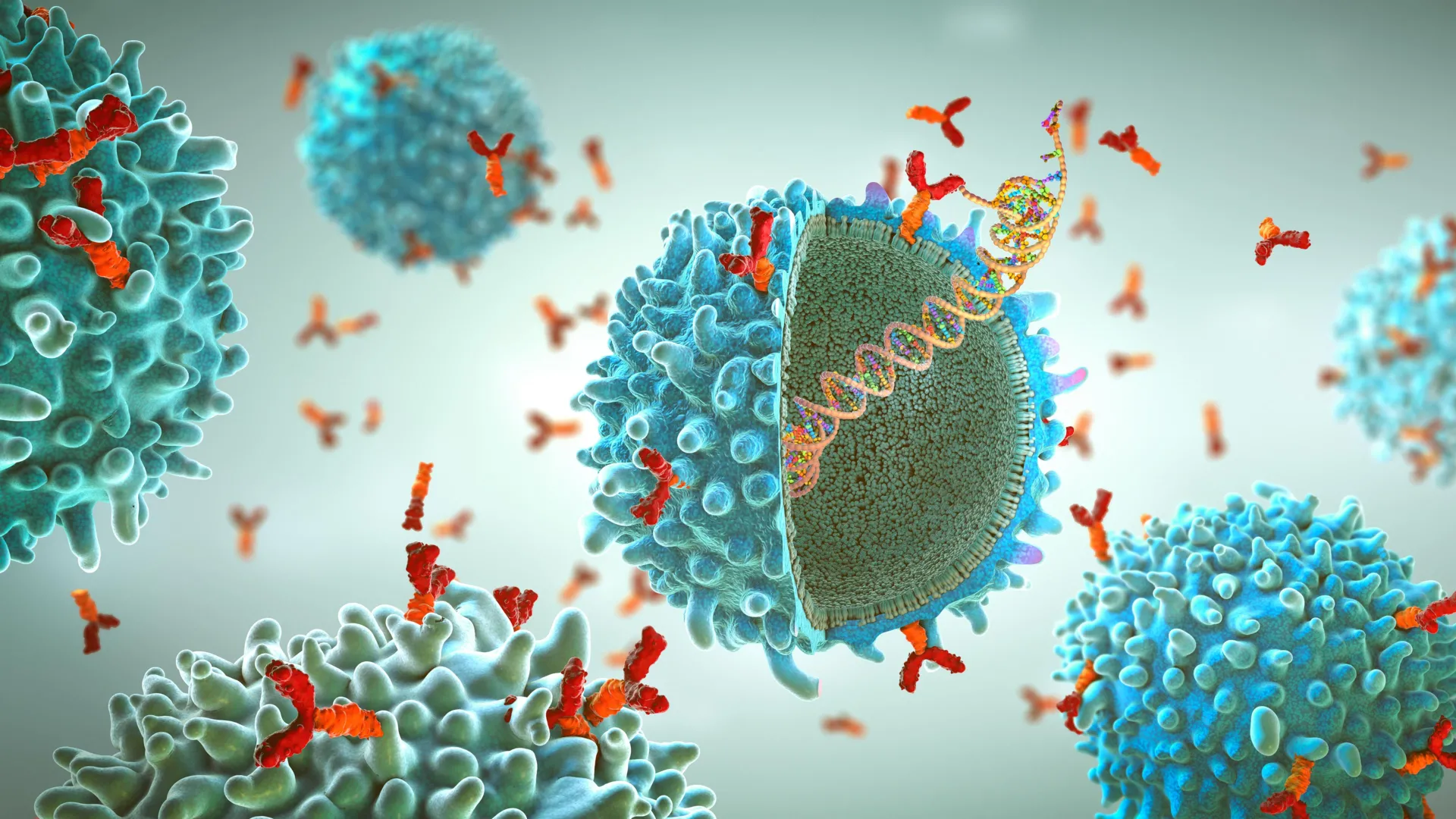
To understand the magnitude of this advancement, one must first understand the limitations of traditional virology. Viruses are characterized by their "spike" or surface glycoproteins, which are anchored into the viral envelope—a lipid membrane that surrounds the virus. These proteins do not exist in isolation; they are held in specific spatial arrangements by the membrane itself.
However, for years, standard laboratory practice has involved "truncating" these proteins. To make them soluble and easier to handle in a petri dish, researchers typically remove the membrane-anchoring segment. While this creates a stable protein for X-ray crystallography or cryo-electron microscopy, it introduces a critical blind spot. Many of the most potent neutralizing antibodies—the "gold standard" for vaccines—target the regions of the protein closest to the membrane. When that anchoring segment is missing, the protein’s shape can shift, and the critical binding sites for these antibodies can be altered, hidden, or completely lost.
"For many years, we’ve had to rely on versions of viral proteins that are missing important pieces," explains Dr. William Schief, co-senior author of the study, a professor at Scripps Research and executive director of vaccine design at IAVI’s Neutralizing Antibody Center. "Our platform lets us study these proteins in a setting that better reflects their natural environment, which is critical if we want to understand how protective antibodies recognize a virus."
Chronology of a Breakthrough: From Concept to Platform
The development of the nanodisc platform was not a sudden epiphany but the culmination of years of iterative research aimed at closing the gap between simplified lab models and the messy reality of biological infection.
- Initial Challenges (Pre-2020): The team recognized that HIV, with its rapidly mutating envelope, was essentially "invisible" to standard vaccine designs. They identified that the base of the protein was a highly conserved, stable region—a "weak point"—that could be exploited if only they could visualize it correctly.
- The Nanodisc Integration: The researchers began experimenting with lipid-based nanodiscs—tiny, self-assembling patches of membrane that can cradle a protein in its natural orientation. By inserting vaccine candidate proteins into these discs, they effectively "re-embedded" the proteins.
- Proof of Concept: The team initially focused on HIV, demonstrating that their nanodisc-bound proteins maintained a native-like architecture. They successfully utilized high-resolution imaging to capture the interaction between these proteins and broad-spectrum neutralizing antibodies.
- Expanding the Scope: Realizing the versatility of the system, the team applied the same methodology to Ebola proteins. The success of this transition proved that the platform was not a "one-off" solution but a scalable, universal tool for virology.
- Current Status: As of the 2024 publication, the platform is being positioned as a standard, high-throughput analytical tool that can accelerate the screening of vaccine candidates, reducing the time required for certain experimental assays from a month to approximately one week.
Supporting Data: Precision and Efficiency
The strength of the Scripps Research platform lies in its marriage of structural accuracy and laboratory efficiency. By incorporating these proteins into nanodiscs, the researchers created a "molecular bait" system. This bait can be used in three distinct, critical ways:
- Structural Analysis: Using advanced imaging, the team gained a level of detail previously inaccessible. They observed how antibodies lock onto the base of the protein at the membrane interface, revealing the precise physical mechanisms required to disrupt viral entry.
- Immune Cell Sorting: The nanodiscs can be used to isolate B-cells—the immune cells responsible for producing antibodies—that specifically target the native-state protein. This allows researchers to "fish out" the most promising candidates from a pool of potential vaccine responders.
- Reproducibility and Scale: First author Kimmo Rantalainen, a senior scientist in the Schief lab, emphasized the importance of the system’s design. "Putting all of these components together into a single, reliable system was the key," Rantalainen noted. "The individual pieces already existed, but making them work together in a way that’s reproducible and scalable opens up new possibilities for how vaccines are analyzed and designed."
The data suggests that this method can significantly improve the "hit rate" for vaccine candidates. By testing them against the nanodisc-bound proteins early in the development cycle, researchers can discard ineffective designs before they reach the costly and time-consuming stage of clinical trials.
Official Responses and Collaborative Impact
The project represents a massive collaborative effort involving experts in structural biology, immunology, and protein engineering. In addition to the lead team at Scripps, the study included contributors from various departments within the institute, as well as industry experts like Sunny Himansu from Moderna Inc.
The funding landscape reflects the high stakes of this research, with significant support from the National Institute of Allergy and Infectious Diseases (NIAID) and the Bill and Melinda Gates Foundation. These organizations have long prioritized the "neutralizing antibody" approach, and the ability to visualize these interactions directly supports their global mission to eradicate diseases that have historically been resistant to vaccine intervention.
The academic community has noted that the platform’s utility extends beyond the pathogens tested. As the researchers pointed out, the structural logic of membrane-bound surface proteins is shared by many of the world’s most dangerous viruses, including influenza and SARS-CoV-2. By providing a "platform" rather than a specific vaccine, the team has effectively gifted the broader scientific community a new lens through which to view viral evolution and immune evasion.
Implications for Future Vaccine Design
While the nanodisc platform is an analytical tool rather than a therapeutic vaccine, its implications for the future of medicine are profound. The history of vaccine development is often characterized by trial and error. Traditional methods involve creating a candidate, testing it, and observing the results—often discovering months later that the protein did not fold correctly or that the immune response was directed at the wrong part of the molecule.
With this new technology, the "design-build-test" cycle becomes significantly more intelligent. Scientists can now:
- Target Conserved Regions: By accurately mapping the membrane-proximal regions of viral proteins, researchers can design vaccines that force the immune system to focus on parts of the virus that cannot mutate easily, potentially leading to more durable, "universal" vaccines.
- Optimize Candidate Selection: The ability to move from a month-long process to a one-week cycle means that labs can screen hundreds of potential designs in the time it previously took to test a handful.
- Enhance Safety and Specificity: By ensuring that the proteins used in vaccines are as close to the "real thing" as possible, researchers can minimize the production of antibodies that are irrelevant or potentially harmful, focusing the body’s energy on neutralizing the virus.
"This gives the field a more realistic, accurate way to test ideas early on," Dr. Schief concluded. "By improving how we study viral proteins and antibody responses, we hope this platform will help advance next-generation vaccines against some of the world’s most challenging viruses."
As the scientific community moves forward, the "Virus glycoprotein nanodisc platform" stands as a testament to the power of fundamental research. By looking closer at the physical interface where a virus meets the world, researchers have found a new way to stop it in its tracks. The journey from the lab bench to the clinic is long, but with this new tool, the path has become significantly clearer, faster, and more precise.